1. Getting started
What is RDF Portal?
RDF Portal is a platform for accessing and integrating life science datasets represented in the Resource Description Framework (RDF). It hosts a wide range of RDF datasets covering genomic, proteomic, chemical, disease, and other biomedical domains.
The portal is maintained by the Database Center for Life Science (DBCLS) and provides multiple ways to access data — from browsing and downloading to querying via SPARQL endpoints and AI-assisted interfaces. For more details on the background, history, and mission of RDF Portal, see the About page.
Key concepts for beginners
If you are new to RDF and SPARQL, here is a brief introduction to the core concepts.
RDF (Resource Description Framework) is a standard model for representing data as a graph. Data is expressed as a collection of statements called triples, each consisting of three parts:
- Subject — the entity being described (e.g., a gene, a protein)
- Predicate — the relationship or property (e.g., “has function”, “is located in”)
- Object — the value or related entity (e.g., a specific function, a chromosome)
Each component is typically identified by a URI (Uniform Resource Identifier), which ensures global uniqueness and allows datasets from different sources to be linked together.
SPARQL is the query language for RDF data. It allows you to retrieve, filter, and combine data across RDF datasets. If you are familiar with SQL for relational databases, SPARQL serves a similar role for graph-based RDF data.
Ontology is a formal representation of knowledge within a domain, defining the types of entities and the relationships between them. RDF Portal datasets use community-standard ontologies such as Gene Ontology (GO), ChEBI, and Disease Ontology to ensure consistent semantics across datasets.
Site navigation
The RDF Portal website is organized into the following main sections, accessible from the left sidebar:

| Section |
Description |
| About |
Background information on RDF Portal, its history, and funding |
| Access methods |
Different ways to query and interact with the data |
| Datasets |
Browsable list of all hosted RDF datasets |
| Statistics |
Summary statistics (triples, classes, properties, etc.) for each dataset |
| Download |
Download RDF data files in various serialization formats |
| Documents |
Manual, data submission guidelines, and RDF config documentation |
| Announcements |
News and updates |
| Update log |
History of data updates |
The site is available in both English and Japanese. You can switch languages using the language link at the bottom of the sidebar.
2. Browsing datasets
Dataset list
The Datasets page displays all RDF datasets hosted on the portal. You can use the controls at the top to sort and filter the list.
Sorting options:
- Date — sort by the dataset registration or update date
- Name — sort alphabetically by dataset name
- Triples — sort by the number of triples (dataset size)
Each sort can be set to descending or ascending order.
Filtering options:
- Tags — filter by domain category (see the tag list below)
- Provenance — filter by data origin
- Registration — filter by how the dataset was registered
Each dataset is assigned one or more tags indicating its domain category. Tags are displayed with petal-shaped icons for easy visual identification.
| Icon |
Tag |
Description |
|
Gene |
Datasets related to genes, gene annotations, and gene-level information |
|
Gene expression |
Datasets containing gene expression profiles and transcriptomics data |
|
Genome |
Datasets related to genome sequences and genomic features |
|
Protein |
Datasets related to protein sequences, structures, and functions |
|
Drug/Chemical |
Datasets related to drugs, chemical compounds, and bioactive molecules |
|
Health/Disease |
Datasets related to diseases, clinical variants, and medical information |
|
Glycan |
Datasets related to glycans and carbohydrate structures |
|
Organism |
Datasets related to organism-level information and taxonomy |
|
Cell |
Datasets related to cell-level information |
|
Bioresource |
Datasets related to biological resource collections (culture collections, biobanks) |
|
Polymorphism |
Datasets related to genetic variants, SNPs, and polymorphisms |
|
Sequence |
Datasets related to nucleotide or amino acid sequences |
A single dataset may have multiple tags. For example, Open TG-GATEs is tagged with Gene, Drug/Chemical, Health/Disease, and Gene expression, reflecting its coverage of toxicogenomics data across these domains.
Provenance
Provenance indicates how the RDF data was created relative to the original data source.
| Value |
Description |
| Original |
RDF data developed by the original database developers themselves. The data provider created the RDF representation of their own data. |
| Third-party |
RDF data developed by a third party, not by the original database developers. Someone other than the original data provider independently converted the publicly available data into RDF. |
Registration
Registration indicates how the dataset was added to RDF Portal.
| Value |
Description |
| Submitted |
The dataset was submitted to RDF Portal by the RDF data developers. |
| Added by RDF Portal |
The dataset was registered by the RDF Portal team. |
Dataset detail page
Clicking on a dataset name opens its detail page. Each detail page contains the following information:
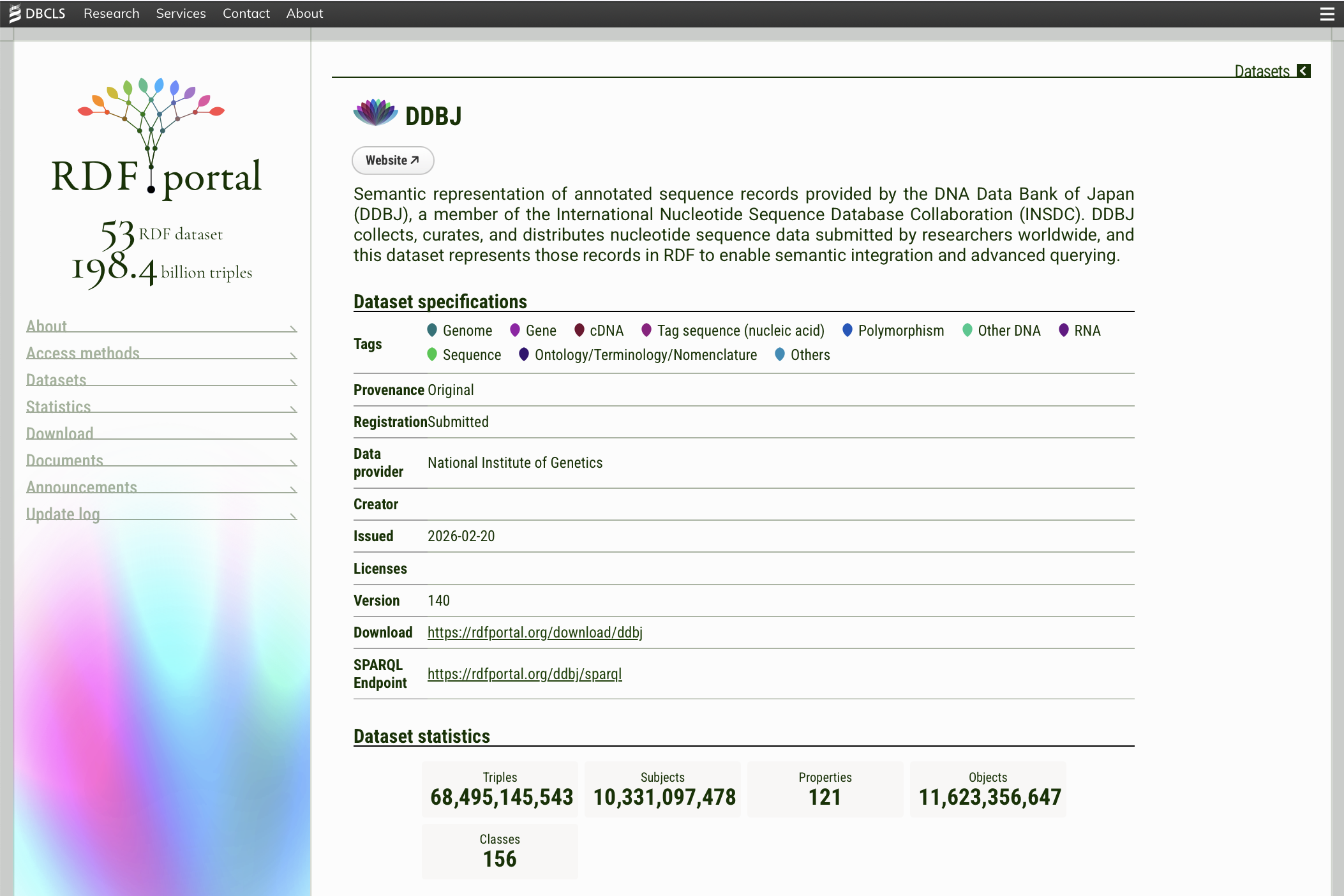
Dataset specifications — a table showing metadata about the dataset:
| Field |
Description |
| Tags |
Domain categories assigned to the dataset |
| Provenance |
Whether the data is original or derived |
| Registration |
How the dataset was added to RDF Portal |
| Data provider |
The organization that provides the data |
| Creator |
The creator of the RDF conversion |
| Issued |
The date the current version was published |
| Licenses |
License information for the dataset |
| Version |
The version number of the dataset |
| Download |
Link to download the RDF data files |
| SPARQL Endpoint |
The SPARQL endpoint URL for querying this dataset |
Dataset statistics — summary counts including the total number of triples, subjects, properties, objects, and classes.
SPARQL example queries — ready-to-use example queries that demonstrate how to retrieve data from the dataset. Each example includes a description and a “Run on Endpoint” link that opens the query directly in the SPARQL endpoint interface.
Schema diagram — a visual representation of the dataset’s RDF schema, showing the classes and properties used in the data. These diagrams are automatically generated from RDF-config, a framework for describing RDF dataset structure in a machine-readable format. RDF-config models are maintained for each dataset hosted on RDF Portal, providing a consistent and practical way to document how the data is organized. In some exceptional cases, schema diagrams may be provided through other means. For more details on RDF-config and its role in RDF Portal, see the RDF config documentation.
3. Access methods
RDF Portal provides several methods for accessing data, ranging from direct SPARQL queries to AI-assisted natural language interfaces.
3a. SPARQL endpoints
SPARQL endpoints allow you to execute SPARQL queries directly against the RDF data. RDF Portal organizes its datasets across multiple SPARQL endpoints, grouped by data source.
For a complete and up-to-date list of available SPARQL endpoints and the datasets they host, see the SPARQL Endpoints page.
Using the SPARQL endpoint in a web browser
Each endpoint provides a web-based query interface. You can access it by visiting the endpoint URL directly (e.g., https://rdfportal.org/ebi/sparql). The interface allows you to:
- Enter a SPARQL query in the text area
- Click “Run” to execute the query
- View the results in tabular format

You can also use the “Run on Endpoint” links provided on each dataset’s detail page to execute the example queries directly.
The web interface is powered by two open-source tools developed by DBCLS:
- SPARQL proxy — A proxy server that sits in front of SPARQL endpoints, providing query validation, job scheduling for concurrent queries, result caching, and logging. It ensures safe and stable access to the endpoints by filtering out potentially harmful queries and managing query workloads.
- Endpoint browser — A web-based interface for browsing and exploring the structure of RDF data stored in SPARQL endpoints. It allows users to visually navigate classes, properties, and their relationships within datasets.
Querying from the command line
You can send SPARQL queries programmatically using tools like curl:
curl -H "Accept: application/sparql-results+json" \
--data-urlencode "query=SELECT ?s ?p ?o WHERE { ?s ?p ?o } LIMIT 10" \
https://rdfportal.org/ebi/sparql
Common Accept header values for specifying the response format:
| Format |
Accept header |
| JSON |
application/sparql-results+json |
| XML |
application/sparql-results+xml |
| CSV |
text/csv |
| TSV |
text/tab-separated-values |
Querying from Python
from SPARQLWrapper import SPARQLWrapper, JSON
sparql = SPARQLWrapper("https://rdfportal.org/ebi/sparql")
sparql.setQuery("""
PREFIX cco: <http://rdf.ebi.ac.uk/terms/chembl#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
SELECT DISTINCT ?molecule_chemblid ?molecule_label
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?Molecule a cco:SmallMolecule ;
cco:chemblId ?molecule_chemblid ;
rdfs:label ?molecule_label .
}
LIMIT 10
""")
sparql.setReturnFormat(JSON)
results = sparql.query().convert()
for result in results["results"]["bindings"]:
print(result["molecule_chemblid"]["value"], result["molecule_label"]["value"])
To install the required library: pip install sparqlwrapper
Querying from R
library(SPARQL)
endpoint <- "https://rdfportal.org/ebi/sparql"
query <- "
PREFIX cco: <http://rdf.ebi.ac.uk/terms/chembl#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
SELECT DISTINCT ?molecule_chemblid ?molecule_label
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?Molecule a cco:SmallMolecule ;
cco:chemblId ?molecule_chemblid ;
rdfs:label ?molecule_label .
}
LIMIT 10
"
results <- SPARQL(endpoint, query)
print(results$results)
Specifying the target graph
Many endpoints host multiple datasets within a single endpoint (e.g., the EBI endpoint hosts ChEBI, ChEMBL, Ensembl, and Reactome). To query a specific dataset, use the FROM clause to specify the named graph:
SELECT ?s ?p ?o
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?s ?p ?o .
}
LIMIT 10
To discover which named graphs are available in an endpoint, you can use the following query:
SELECT DISTINCT ?g
WHERE {
GRAPH ?g { ?s ?p ?o }
}
3b. GraphQL API
This section will be available once the GraphQL API documentation is published.
The GraphQL API provides an alternative query interface designed for application developers. It allows schema-based, intuitive query construction and is particularly suitable for frontend application integration.
3c. MCP interface (TogoMCP)
TogoMCP is a Model Context Protocol (MCP) interface for AI agents. It enables large language models (LLMs) to understand the structure and content of RDF datasets, allowing them to retrieve accurate data in response to natural language questions.
The TogoMCP server URL is: https://togomcp.rdfportal.org/mcp
For detailed documentation on how to set up and use TogoMCP, including configuration guides and usage examples, please refer to the TogoMCP website.
3d. LLM chat interface
TBA
3e. SPARQL composer
The SPARQL composer is a tool for generating SPARQL queries through a graphical interface. It is useful for users who want to construct queries visually without writing SPARQL syntax manually.
4. Statistics
The Statistics page provides a summary table showing key metrics for each dataset hosted on RDF Portal.

Understanding the statistics table
| Column |
Description |
| Dataset |
The name of the dataset (links to the dataset detail page) |
| Triples |
The total number of RDF triples in the dataset. This is the primary measure of dataset size. |
| Classes |
The number of distinct RDF classes (types of entities) defined or used in the dataset. |
| Properties |
The number of distinct predicates (relationships) used in the dataset. |
| Subjects |
The number of distinct subject URIs. This roughly corresponds to the number of unique entities described in the dataset. |
| Objects |
The number of distinct object values (both URIs and literals). |
Interpreting the numbers
The datasets on RDF Portal vary enormously in size. For example:
- DDBJ is the largest dataset with approximately 68.5 billion triples, containing nucleotide sequence data from the DNA Data Bank of Japan.
- UniProt RDF contains over 51.3 billion triples of protein sequence and functional information.
- Nucleic Acid Drug Database is one of the smallest datasets with 948 triples.
The ratio of classes to properties gives an indication of the dataset’s schema complexity. A dataset with many classes and properties (e.g., wwPDB/RDF with 647 classes and 3,823 properties) has a rich, detailed data model, while a dataset with few classes and properties (e.g., PubMed with 2 classes and 5 properties) has a simpler, flatter structure.
5. Download
The Download page provides links to download RDF data files for each dataset. Data is available in multiple RDF serialization formats.
| Format |
Extension |
Description |
Best for |
| N-Triples |
.nt |
One triple per line, simple text format |
Streaming, bulk loading, line-by-line processing |
| Turtle |
.ttl |
Compact, human-readable format with prefix abbreviations |
Manual inspection, readability |
| RDF-XML |
.rdf |
XML-based serialization |
XML toolchains, legacy systems |
| JSON-LD |
.jsonld |
JSON-based linked data format |
Web applications, JavaScript environments |
Not all formats are available for every dataset. The availability follows these rules:
- Original submitted files — Every dataset provides at minimum the RDF files in the format originally submitted by the data provider. This is always available.
- N-Triples — In addition to the original format, an N-Triples (
.nt) version is always generated and provided for every dataset. N-Triples serves as the common baseline format, ensuring that all datasets can be processed uniformly regardless of the original submission format.
- Other formats (Turtle, RDF-XML, JSON-LD) — These are provided when available, but are not guaranteed for every dataset. Availability depends on whether the conversion has been performed for that particular dataset.
On the Download page, a dash (—) in a format column indicates that the format is not currently available for that dataset.
- For bulk loading into a triplestore (e.g., Virtuoso, GraphDB, Apache Jena), N-Triples is generally the fastest format to parse and is always available.
- For reading and understanding the data structure, Turtle provides the most human-friendly representation.
- For web application integration, JSON-LD is the natural choice as it can be processed directly by JavaScript.
- For compatibility with XML-based tools, RDF-XML is appropriate.
Download URLs
Download links follow the pattern: https://rdfportal.org/download/{dataset_id}
For example, to download ChEMBL RDF data: https://rdfportal.org/download/chembl
6. Use cases and tutorials
This section provides practical examples of how to use RDF Portal data for life science research.
Tutorial 1: Your first SPARQL query
This tutorial walks you through executing a simple SPARQL query to retrieve data from ChEMBL.
Goal: List 10 small molecule compounds with their ChEMBL IDs and names.
Step 1: Open the EBI SPARQL endpoint at https://rdfportal.org/ebi/sparql
Step 2: Enter the following query:
PREFIX cco: <http://rdf.ebi.ac.uk/terms/chembl#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
SELECT DISTINCT ?molecule_chemblid ?molecule_label
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?Molecule a cco:SmallMolecule ;
cco:chemblId ?molecule_chemblid ;
rdfs:label ?molecule_label .
}
LIMIT 10
Step 3: Click “Run” to execute the query. You will see a table of results showing ChEMBL IDs and molecule names.
Understanding the query:
PREFIX lines define namespace abbreviations used in the querySELECT specifies which variables to returnFROM specifies the named graph (dataset) to queryWHERE defines the pattern to match — here, we look for entities that are typed as SmallMolecule and have both a ChEMBL ID and a labelLIMIT 10 restricts the output to 10 results
Tutorial 2: Finding approved drugs for a specific target
Goal: Find compounds approved as drugs (development phase 4) that target Tyrosine-protein kinase ABL.
PREFIX cco: <http://rdf.ebi.ac.uk/terms/chembl#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX chembl_target: <http://rdf.ebi.ac.uk/resource/chembl/target/>
SELECT ?molecule_chemblid ?molecule_label
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?Molecule a cco:SmallMolecule ;
cco:chemblId ?molecule_chemblid ;
rdfs:label ?molecule_label ;
cco:highestDevelopmentPhase 4 ;
cco:hasMechanism ?mechanism .
?mechanism cco:hasTarget chembl_target:CHEMBL1862 .
}
LIMIT 100
This query combines multiple conditions: filtering by molecule type, development phase, and a specific drug target. Modify chembl_target:CHEMBL1862 to search for drugs targeting other proteins.
Tutorial 3: Cross-endpoint federated queries
SPARQL supports federated queries using the SERVICE keyword, which allows you to combine data from multiple endpoints in a single query.
Goal: Retrieve UniProt protein entries and link them to their corresponding Reactome pathways.
PREFIX up: <http://purl.uniprot.org/core/>
PREFIX biopax3: <http://www.biopax.org/release/biopax-level3.owl#>
SELECT ?protein ?proteinName ?pathway ?pathwayName
WHERE {
SERVICE <https://rdfportal.org/sib/sparql> {
?protein a up:Protein ;
up:mnemonic ?proteinName .
FILTER(CONTAINS(?proteinName, "HUMAN"))
}
SERVICE <https://rdfportal.org/ebi/sparql> {
?pathway a biopax3:Pathway ;
biopax3:displayName ?pathwayName .
}
}
LIMIT 10
Note: Federated queries can be slow depending on the size of the intermediate results. Always use LIMIT and apply FILTER conditions to reduce the data transferred between endpoints.
Tutorial 4: Exploring dataset structure
Before writing queries for a new dataset, it is helpful to explore its structure. The following queries can be used with any endpoint.
List all classes in a dataset:
SELECT DISTINCT ?class (COUNT(?s) AS ?count)
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?s a ?class .
}
GROUP BY ?class
ORDER BY DESC(?count)
List all properties used in a dataset:
SELECT DISTINCT ?property (COUNT(?s) AS ?count)
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?s ?property ?o .
}
GROUP BY ?property
ORDER BY DESC(?count)
Get sample data for a specific class:
PREFIX cco: <http://rdf.ebi.ac.uk/terms/chembl#>
SELECT ?s ?p ?o
FROM <http://rdf.ebi.ac.uk/dataset/chembl>
WHERE {
?s a cco:SmallMolecule ;
?p ?o .
}
LIMIT 20
7. FAQ and troubleshooting
General questions
Q: Is RDF Portal free to use?
A: Yes. RDF Portal is a publicly funded infrastructure and all data access is free of charge. Individual datasets may have their own licenses — check the “Licenses” field on each dataset’s detail page.
Q: How often is the data updated?
A: Update frequency varies by dataset. Check the Update log page for the latest update history.
Q: Can I submit my own RDF dataset?
A: Yes. Please refer to the Data submission guidelines. All submitted datasets undergo a quality review by DBCLS to ensure compliance with the DBCLS RDF Guidelines.
SPARQL query issues
Q: My query is timing out. What should I do?
A: Try the following approaches:
- Add a
LIMIT clause to restrict the number of results
- Use more specific
FILTER conditions to narrow the search
- Avoid
SELECT * — specify only the variables you need
- Use
FROM to target a specific named graph rather than querying the entire endpoint
- For very large result sets, consider downloading the data files and loading them into a local triplestore
Q: How do I know which named graph to use in the FROM clause?
A: Each dataset’s detail page shows the SPARQL endpoint URL and named graph URI. You can also discover named graphs by running:
SELECT DISTINCT ?g WHERE { GRAPH ?g { ?s ?p ?o } }
Q: My query returns no results. What could be wrong?
A: Common causes include:
- Incorrect namespace URIs — check the prefix declarations against the dataset’s schema diagram
- Wrong named graph — verify the
FROM clause matches the dataset’s graph URI
- Case sensitivity — URIs and literal values are case-sensitive in SPARQL
- Data type mismatches — when filtering by numeric values, ensure the type matches (e.g., integer vs. string)
Data and licensing
Q: Can I redistribute the data I download?
A: This depends on the license of each individual dataset. Check the “Licenses” field on the dataset’s detail page. Many datasets are available under open licenses that permit redistribution.
Q: How should I cite RDF Portal?
A: Please cite the RDF Portal website URL (https://rdfportal.org/) and the specific dataset(s) you used. For individual datasets, cite the original data providers as specified on each dataset’s detail page.
Q: How should I cite RDF Portal?
A: Please cite the following publication:
Shuichi Kawashima, Toshiaki Katayama, Hideki Hatanaka, Tatsuya Kushida, Toshihisa Takagi. NBDC RDF portal: a comprehensive repository for semantic data in life sciences. Database (Oxford). 2018 Jan 1;2018:bay123. doi: 10.1093/database/bay123. PubMed: 30576482.
For questions, bug reports, or feedback, please contact the DBCLS team through the information provided on the About page.